
Molecular diagnostics comprises a rapidly evolving range of assays for the detection and analysis of nucleic acid sequences and proteins. These enable the detection of pathogens, estimation of viral loads; help with the selection of antibiotic and antiviral therapies, diagnosis of cancer and other diseases and offer prognostic assessments as well as assistance with treatment selection and drug treatment efficacy monitoring. Inappropriate use of molecular technologies is also leading to serious issues with the clinical relevance of many molecular biomarkers in use today. There is an urgent need for standardisation of assay designs, laboratory practices, measurement methods and data management.
Molecular diagnostics is becoming increasingly important as an essential tool for the rapid, accurate and cost-effective delivery of safe and effective therapy for many diseases. Its uptake is driven by (i) continuous innovation in underlying technologies, (ii) constant identification of new or improved biomarkers, (iii) steady decrease in associated costs and (iv) increasing automation that enables even complex molecular assays to be performed in a wide range of health care settings. Tests may be relatively unsophisticated, such as those used for the detection of pathogen-specific genomic sequences; they may involve the identification of complex disease-associated mutational and epigenetic changes in DNA sequences or differential expression patterns of RNA and proteins or they may allow the identification of a predisposition to a particular disease or screen for diseases that are present but asymptomatic. The information provided by genomic sequencing is critical for our understanding of the interactions that determine an individual’s health at a molecular level and whereas the first draft sequence of the human genome took ten years of world-wide effort and cost in the region of US$2.7 billion, it is now possible to sequence a complete human genome in less than a day at a cost of around US$1,000. This is beginning to allow the identification of every individual's unique molecular characteristics and to apply them to personalised diagnosis, treatment and susceptibility prediction
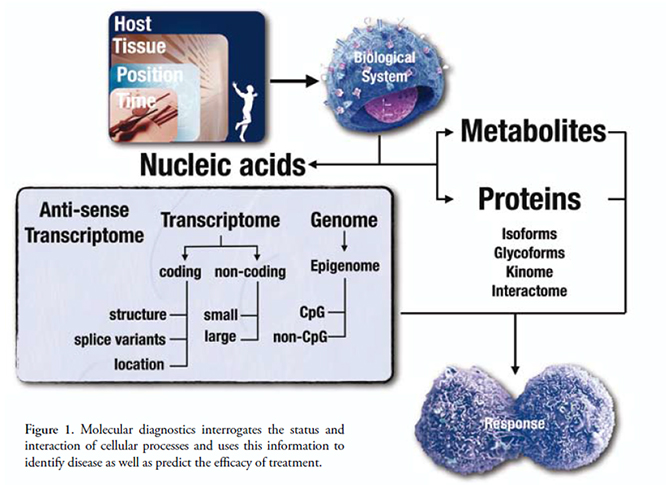
However, it is essential that laboratory methods and procedures are fully validated and verified before their use in clinical testing to ensure that assays are carried to clear standards. Biomarkers need to be chosen carefully to understand the complex relationship between predictive and prognostic biomarkers. Furthermore, technological advance has been such that the ability to collect and analyse data has become a major bottleneck, especially with data generated by Next Generation Sequencing (NGS) methods. This requires the development of integrated and dynamic data collection methods that are standardized between science and healthcare. Finally, the regulation of genetic testing, together with ethical considerations within an appropriate legal framework constitutes to challenge society as a whole and must be resolved to ensure the proper future of molecular diagnostics.
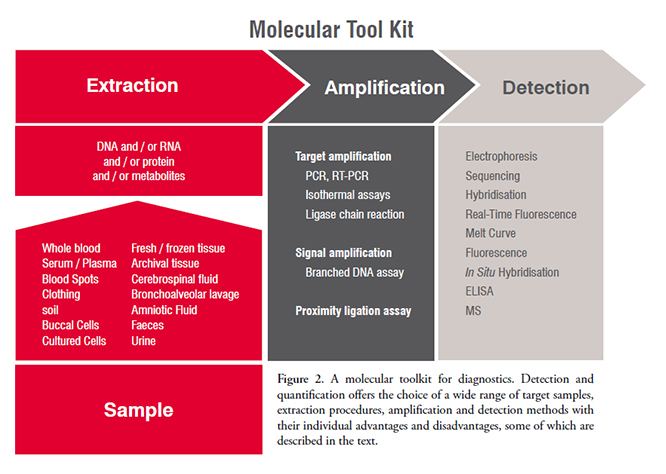
Molecular diagnostics comprises a huge range of technologies ranging from comparatively simple techniques such as the real-time Polymerase Chain Reaction (qPCR) for the quantification of nucleic acids to complex technologies such as Mass Spectrometry (MS), which analyses protein-based biomarkers and NGS, which allows the definitive identification of specific nucleic acid sequences. Most assays follow a general workflow that involves some kind of extraction, amplification and final detection steps (Figure 2). In general, Nucleic Acid-based Tests (NATs) are most widely used and encompass a range of amplification technologies and sophisticated detection methods that are readily adaptable for use in the diagnostic laboratory. However, reliance on knowledge of the nucleic acid sequence introduces an important limitation of most NATs (NGS is the exception). Assay design requires comprehensive information on sequences and sequence variations for targeted cellular targets or pathogens. In their absence, most NATs cannot detect unknown mutations or pathogens. However, as with the human genome, the availability of genome sequences for more than 1,000 bacterial, 3,000 viral and nearly 100 fungal genomes, including representatives of all significant human pathogens, has revolutionised the fields of microbiology and infectious diseases. It permits the increasing penetration of NATs into pathogen diagnosis and genotyping as well as the detection of novel virulence and antibiotic resistance markers. These developments are entrenching NATs as the benchmark for diagnosis of an ever-increasing range of pathogens, and their use is driving improvements in disease management whilst helping to decrease the costs associated with patient care.
PCR is the most widely used technology in molecular diagnostics, with other techniques such as NGS often relying on its powerful amplification ability to generate target molecules for further investigation. Other NATs are used in molecular diagnostics and their characteristics are discussed elsewhere. Today qPCR is at the heart of numerous diagnostic assays, with its potential for quantification enabling assessment of DNA copy number variation, differential RNA expression levels and pathogen load. It can even be combined with disease- or pathogen-specific antibodies in a method termed ‘proximity ligation assay’, an innovative immunoassay platform that combines the exquisite sensitivity and dynamic range of qPCR with antibody-based detection of proteins and other analytes to quantify protein levels, providing a more sensitive alternative to conventional ELISAs.
Whereas the original PCR protocols assessed individual markers, the trend now is towards testing for multiple targets in a single assay (“multiplex”), which not only allows for high throughput testing but also reduces cost and the time taken to complete the assay:
HPV testing provides an excellent example of the usefulness of PCR-based testing for pathogen screening, which is used to identify HPV subtypes associated with an increased risk of cervical cancer
Cystic Fibrosis (CF) is a disease with numerous common mutations in the CF Transmembrane Regulator (CFTR) gene and multiplex PCR is an example of a screening/diagnostic test that helps identify carriers or can be used for foetal CF assessment
Detection of KRAS mutations is an example of a molecular diagnostic test that helps with personalised therapeutic decision-making, since tumours harbouring mutations in this gene fail to respond to drugs such as Erbitux and Vectibix that target the Epidermal Growth Factor Receptor (EGFR).
Despite its ubiquity, there are several weaknesses to PCR in general and qPCR in particular. First, a high degree of performance variability has been observed between different DNA tests due to poor standardisation of PCR assay designs. Second, the requirement for purified nucleic acids limits the practicality of qPCR in a clinical setting, since it increases the time needed for sample preparation and introduces the likelihood of contamination. qPCR is also sensitive to environmental inhibitors that are concentrated along with target nucleic during sample processing. However, Cepheid’s GeneXpert technology, which uses an integrated microfluidic-based cartridge system to automatically extract nucleic acids and carry out multiplex qPCR assays points the way to how these problems are being overcome.
Third, especially important for pathogen detection, the increasingly small volumes assayed may lead to false-negative results because of low pathogen counts or titers. Consequently, for many diagnostic applications there is no advantage to carrying out PCR reactions in nanolitre volumes, which has the benefit of making the assay workflow less specialised and reliant on expensive instrumentation.
Fourth, since PCR only detects nucleic acids, a positive result offers no conclusive evidence of pathogen viability or active growth. Hence the detection of an organism’s DNA in a sample does not directly indicate the presence of an active infection and, therefore, can lead to a false-positive result. This is a problem with infectious aspergillosis, where early initiation of systemic antifungal therapy is compromised by a lack of definitive diagnostic criteria and antibody detection lacks the ultimate sensitivity required to detect pathogens at the earliest stages of infection. Moreover, DNA tests are highly susceptible to environmental contamination, since dormant Aspergillus spores are ubiquitous. PLA may offer a solution to this challenge, as it offers several advantages over traditional ELISAs, including 10 to 100-fold better sensitivity, broader dynamic range, simpler workflow and faster time to results. By using PLA to detect quantitatively proteins expressed exclusively by actively growing fungi, this assay could provide early confirmation of active infection.
A major problem with all PCR-based assays is that the basic protocols, whilst superficially simple, are persistently poorly performed. The most egregious example of the misuse of PCR technology is illustrated by the publication of seriously flawed data purporting to show the presence of measles virus in the intestinal epithelium of autistic children 30. Three recent surveys of the qPCR-based literature suggest that a very high percentage of these publications are likely to contain biologically or clinically meaningless results, a situation that probably explains the many discordant results found in the peer-reviewed literature. This is due to variable pre-assay conditions, absent quality controls, poor assay design and incorrect data analysis. In addition, the problem of published data that are often inconsistent, inaccurate and wrong is exacerbated by a lack of transparency of reporting, with the details of technical information inadequate for the purpose of assessing the validity of published qPCR data. Undoubtedly this is in part responsible for the lack of progress in many areas that rely on accurate analysis of gene expression profiles and the resultant translation to clinical tests.
The MIQE (minimum information for the publication of quantitative real-time PCR) guidelines were published in 2009 with the twin aims of providing a blueprint for good real-time quantitative Polymerase Chain Reaction (qPCR) assay design and encouraging the comprehensive reporting of qPCR protocols. MIQE represents a major milestone in the transformation of the qPCR into a reliable technology fit for purpose as a diagnostic technique and has been augmented by complementary guidelines for digital PCR. Unfortunately, inadequate qPCR-based publications continue to appear, adding to the vast numbers already published, which report contradictory and not reproducible data, without any obvious criteria to distinguish a genuine result from a technical artifact.
Microchips, often referred to as “biochips”, are arrays of immobilised biomolecules that allow rapid analysis and sequencing of DNA and detection of RNA as well as of proteins. For DNA microarrays, thousands of spots are arrayed in orderly rows and columns on a solid surface (usually glass). Each spot contains multiple identical strands of DNA, with the DNA sequence on each spot being unique and representing one gene.The precise location and sequence of each spot is recorded in a computer database. In addition to the biochips, this technology requires instruments to handle the samples, read the reported molecules and analyse the resulting wealth of data. They allow massively parallel analysis of many target molecules in many samples, allowing the identification of disease-associated mutations or mRNA/miRNA expression patterns.
In general, DNA-targeting approaches are sufficiently robust and reliable for the American College of Medical Genetics (ACMG) to recommend the use of microarrays as the first-tier diagnostic test in postnatal cytogenetics for detecting Copy Number Variants (CNVs) associated with intellectual disability, developmental delay, and dysmorphicfeatures.Illumina have developed their MiSeqDx Cystic Fibrosis System, which can detect 139 clinically relevant CFTR variants and identifies all variants in the protein coding regions and intron/exon boundaries of CFTR.
In contrast, although cDNA microarray-based results are claimed to be comparable across multiple laboratories, problems of platform compatibility, standardisation and data analysis remain and have contributed to the publication of numerous discordant results.
Whole transcriptome shotgun sequencing (“RNA-Seq”), which offers an alternative approach for high-throughput transcriptome analysis and is used to calculate the prevalence of transcripts, provides new possibilities for biomarker identification. However, it relies on the enzymatic conversion of RNA into DNA (‘reverse transcription’), which is known to be variable and can distort quantitative results. Reliable application of this technology will require not just improvements in sample preparation and sequencing methods, but a more thorough understanding of the limitations of the RT step in quantitative analysis. In addition, clinical NGS testing will place significant demands on laboratory infrastructure, as it requires investment in extensive instrumentation as well as bioinformatics expertise if it is to transfer successfully into clinical practice.
MS is used in molecular diagnostics as a high specificity tool for identifying and quantifying disease associated molecules and proteins. There are different types of MS in use, most commonly Liquid Chromatography (LC-MS), Gas Chromatography (GC-MS), and Matrix-Assisted Laser Desorption/Ionization/Time-Of-Flight (MALDI-TOF MS). MS technology can be used for a wide range of applications that include diagnosis of viral infections, disease screening, diagnosis, detection of metabolic disorders and therapeutic monitoring. The main challenges to a more widespread adoption of this technology lie in complex sample preparation procedures, instrument-specific molecule databases, laboratory information system interfacing and inter-instruments standardisation.
FISH is a well-established technique that has traditionally been used to detect specific DNA sequences on chromosomes and is now widely used for the diagnosis, prognosis and therapeutic monitoring of cancer. One of the most exciting new developments in FISH is termed Stellaris, which allows simultaneous detection, localisation and quantification of individual RNA molecules at the sub-cellular level in fixed samples using wide field fluorescence microscopy. A set of Stellaris FISH Probes comprises multiple oligonucleotides with different sequences each with a fluorescent label that collectively bind along the same target transcript to produce a punctate signal. The Stellaris technology uses a simple protocol with standard reagents and is inexpensive and platform-independent. Its ability to quantify accurately very low copy number targets, together with improvements to the hybridization protocols that speed up the time taken to complete the assay, makes it a useful candidate for the detection of disease-associated transcripts and splice variants, for example those expressed by micrometastases.
Increased throughput, development of nanotechnology and the introduction of Point of Care (POC) devices are confirming molecular diagnostics as a transformative procedure. With molecular techniques increasingly moving from research laboratories into mainstream clinical applications, they are being translated into methods for helping health workers improve clinical outcomes for their patients. This process is still very much in its infancy and the combination of increased understanding of the biological mechanisms underlying disease with the rapid evolution of molecular technologies holds out much promise for the future of molecular diagnostics. The future is likely to see the establishment of new disease/gene associations, widespread use of personalised therapy and the introduction of predictive tests, opening the path to prevention of some diseases. However, accurate diagnostics requires the validation of clinically relevant biomarkers, sound methodologies, reliable methods and transparent reporting of research results and clinical trials. Unfortunately, this is an area that poses significant challenges, but is widely ignored. Finally, there remain concerns about regulatory and ethical considerations being able to keep up with the relentless pace of innovation, requiring informed input from a wide range of stakeholders, most of which have only the vaguest understanding of the challenges associated with this area.